Universal RNA Kit
Kit for extraction and purification of total RNA from human and animal tissues, cell culture, plant, yeast and bacteria.
Detailed Product Description
English Version
 Detailed Reference Manual (English Version Only)
Detailed Reference Manual (English Version Only)
No DNase Digestion Required
The Universal RNA Kit allows to prepare high-quality DNA-free total RNA
- without any DNase digestion step and
- without any organic solvents such as phenol / chloroform.
For virtually all applications, DNase digestion is not required, thus saving lab time, material and eleminating any unwanted RNA loss due to residual RNase activity, which remains present in many "RNase free" DNase preparations.
In most applications, purified RNA is of considerably higher quality as compared to RNA obtained from "classic" phenol / chloroform based extraction methods. The RNA purification procedure is extremely gentle: RNA is neither degraded by residual RNase activity present in DNase preparations, nor affected by the attack of free radicals present in phenol formulations.
Checking DNA-Free Total RNA by Quality Control PCR
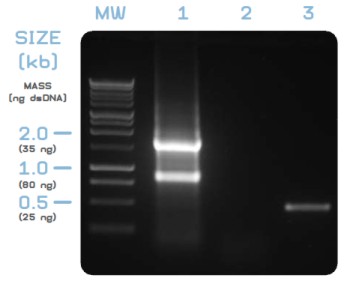
Figure 1: Quick quality control of purified RNA (GeneMatrix Universal RNA Kit), isolated from human dermal fibroblasts, on an 1% [w/v] agarose gel.
- Lane MW: PerfectPlus 1 kb Ladder [5 µl].
- Lane 1: Purified, non-DNase treated RNA from human dermal fibroblasts. 3 μl purified RNA from a total elution volume of 50 μl were directly applied on the gel. 28S / 18S rRNA ratio is visibly >2, indicating high quality of purified RNA. No band at ~20 kb is observed, indicating purified RNA is free from genomic DNA contamination.
- Lane 2: PCR from purified, non-DNase digested RNA did not yield any PCR product, confirming the absence of contaminating DNA from purified RNA.
- Lane 3: PCR from cDNA (derived from the RNA sample shown in lane 2), specific for the human β-actin "housekeeping" gene as described below, gave an amplicon of the expected size (470 bp). No band with a size of 565 bp is present, indicating cDNA was entirely derived from RNA and did not contain contaminating traces of genomic DNA.
- PCR and RT-PCR conditions: RNA was extracted from human dermal fibroblasts following the kit supplied protocol for cell culture RNA isolation. DNase digestion was not performed. Oligo dT primed cDNA was synthesized from purified RNA using EURx AMV Reverse Transcriptase Native, as described in the product manual. Human β-actin gene specific PCR from purified RNA and from cDNA was conducted using OptiTaq DNA Polymerase with the PCR conditions outlined below. 3 µl each of purified RNA and of both PCR reactions, respectively, were analyzed on an 1% [w/v] agarose gel.
- Note: Due to general differences in form factors, secondary structure formation and resulting migration behaviour between single- and double-stranded nucleic acids, 28S rRNA (~5030 nt) and 18S rRNA (~1900 nt) migrate faster through the non-denaturing agarose gel matrix as compared to dsDNA MW marker bands.
Total RNA Quality Control PCR Parameters
Purity of RNA from DNA contamination is experimentally confirmed by PCR. For purified RNA, quality control PCR is conducted as follows: (1) 1 µl of purified RNA and (2) 1 µl of genomic DNA or cDNA (serving as positive control), respectively, are used as templates for setting up two PCR reactions.
Quality control PCR conditions for RNA from human sources: Gene-specific primers for the "housekeeping" human beta-actin gene ß-Ac-fwd (5‘-CGG ATG TCC ACG TCA CAC TT-3‘), ß-Ac-rev (5‘-GTT GCT ATC CAG GCT GTG CT-3‘). PCR amplification parameters 95°C/120 s / 30x [94°C/30s – 60°C/30s – 72°C/60 s] / 72°C/600 s, as described elsewhere (Adjaye et al. (1999) Gene 237: 373–383). RNA quality control PCRs from non-human sources require other, organism-specific primers and PCR conditions.
Evaluation:
- PCR starting from purified total RNA as template must not yield any product. PCR must remain negative to ensure that RNA is DNA-free.
- PCR starting from human genomic template DNA yields a ~ 565 bp amplicon.
- PCR starting from human cDNA templates yields a ~ 470 bp amplicon.
Assessing the Physical Integrity of Purified RNA
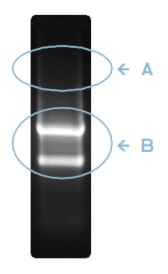
Figure 2: Integrity check of purified RNA, as judged by quick, non-laborious agarose gel electrophoresis. 2.5 - 5 µl (~5 to 10%) of purified RNA is checked on an 1% [w/v] agarose gel. Although there exist certain organism- and tissue-specific patterns of RNA expression, these two common criteria indicate high quality RNA:
(A) High quality RNA does not contain DNA contaminations (observable as ~20 kb band in DNA-contaminated RNA preparations; to be independently verified by direct PCR from purified RNA, which must remain negative).
(B) In high quality RNA preparations, ribosomal RNAs, in particular the 28S rRNA (~5030 nt) and 18S rRNA (~1900 nt) are present at a 1:1 stochiometric ratio and form two prominently visible bands (migrating considerably faster as compared to dsDNA markers). As theoretically inferred from molecule sizes, the 28S rRNA band should appear 2.7 times as bright as the 18S rRNA band. Considering sample specific variations, an 28S to 18S ratio of >2 is indicative for high quality RNA. 28S/18S ratios <2 indicate RNA degradation and are commonly observed in poor quality RNA preparations.
For fully completing quality control, check RNA purity by spectrophotometric measurement in parallel.
Protocols available for RNA extraction from
- human and animal tissue samples;
- plant tissue samples;
- bacterial cells;
- yeast cells;
- cell culture (human and animal cell lines);
- RNA clean-up from molecular biological reactions
(such as T7 mediated RNA transcription). - Additional protocols:
- Optional proteinase K digestion step (solubilization of refractory tissue samples);
- optional organic solvent (phenol/chloroform) digestion step (disintegration of otherwise imperishable samples, such as heart valves);
- optional on-column DNase digestion step (not required for almost all applications).
- dART cDNA Synthesis Kit with all required and specifically adjusted components for Reverse Transcription
- Reverse Transcriptases
- Ribonuclease Inhibitor
- Ribonuclease H ( E.coli ) for selective RNA digestion from DNA/RNA hybrids only
- SG qPCR Master Mixes (2x) for qPCR / RealTime PCR
- T7 Transcription Kit
Additional Resources
 RT Logsheet
RT Logsheet qPCR Logsheet
qPCR Logsheet
